 https://doi.org/10.4081/ejtm.2023.11553
https://doi.org/10.4081/ejtm.2023.11553
Mass spectrometry-based proteomic characterization of the middle-aged mouse brain for animal model research of neuromuscular diseases
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Published: 3 August 2023
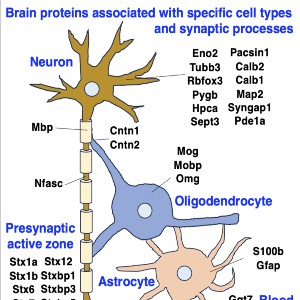
Neuromuscular diseases with primary muscle wasting symptoms may also display multi-systemic changes in the body and exhibit secondary pathophysiological alterations in various non-muscle tissues. In some cases, this includes proteome-wide alterations and/or adaptations in the central nervous system. Thus, in order to provide an improved bioanalytical basis for the comprehensive evaluation of animal models that are routinely used in muscle research, this report describes the mass spectrometry-based proteomic characterization of the mouse brain. Crude tissue extracts were examined by bottom-up proteomics and detected 4558 distinct protein species. The detailed analysis of the brain proteome revealed the presence of abundant cellular proteoforms in the neuronal cytoskeleton, as well as various brain region enriched proteins, including markers of the cerebral cortex, cerebellum, hippocampus and the olfactory bulb. Neuroproteomic markers of specific cell types in the brain were identified in association with various types of neurons and glia cells. Markers of subcellular structures were established for the plasmalemma, nucleus, endoplasmic reticulum, mitochondria and other crucial organelles, as well as synaptic components that are involved in presynaptic vesicle docking, neurotransmitter release and synapse remodelling.
Downloads
Navabpour S, Kwapis JL, Jarome TJ. A neuroscientist's guide to transgenic mice and other genetic tools. Neurosci Biobehav Rev. 2020 Jan;108:732-748. Epub 2019 Dec 13. PMID: 31843544; PMCID: PMC8049509. DOI: https://doi.org/10.1016/j.neubiorev.2019.12.013
Partridge TA. The mdx mouse model as a surrogate for Duchenne muscular dystrophy. FEBS J. 2013 Sep;280(17):4177-86. Epub 2013 Apr 22. PMID: 23551987; PMCID: PMC4147949. DOI: https://doi.org/10.1111/febs.12267
Schmitt-John T. VPS54 and the wobbler mouse. Front Neurosci. 2015 Oct 21;9:381. PMID: 26539077; PMCID: PMC4612502. DOI: https://doi.org/10.3389/fnins.2015.00381
Brown RH, Al-Chalabi A. Amyotrophic Lateral Sclerosis. N Engl J Med. 2017 Jul 13;377(2):162-172. PMID: 28700839. DOI: https://doi.org/10.1056/NEJMra1603471
Akçimen F, Lopez ER, Landers JE, Nath A, Chiò A, Chia R, Traynor BJ. Amyotrophic lateral sclerosis: translating genetic discoveries into therapies. Nat Rev Genet. 2023 Apr 6. Epub ahead of print. PMID: 37024676. DOI: https://doi.org/10.1038/s41576-023-00592-y
Capitanio D, Moriggi M, Gelfi C. Mapping the human skeletal muscle proteome: progress and potential. Expert Rev Proteomics. 2017 Sep;14(9):825-839. Epub 2017 Aug 14. PMID: 28780899. DOI: https://doi.org/10.1080/14789450.2017.1364996
Sharma K, Schmitt S, Bergner CG, Tyanova S, Kannaiyan N, Manrique-Hoyos N, Kongi K, Cantuti L, Hanisch UK, Philips MA, Rossner MJ, Mann M, Simons M. Cell type- and brain region-resolved mouse brain proteome. Nat Neurosci. 2015 Dec;18(12):1819-31. Epub 2015 Nov 2. PMID: 26523646; PMCID: PMC7116867. DOI: https://doi.org/10.1038/nn.4160
Duan D, Goemans N, Takeda S, Mercuri E, Aartsma-Rus A. Duchenne muscular dystrophy. Nat Rev Dis Primers. 2021 Feb 18;7(1):13. PMID: 33602943. DOI: https://doi.org/10.1038/s41572-021-00248-3
Ohlendieck K, Swandulla D. Complexity of skeletal muscle degeneration: multi-systems pathophysiology and organ crosstalk in dystrophinopathy. Pflugers Arch. 2021 Dec;473(12):1813-1839. Epub 2021 Sep 22. PMID: 34553265; PMCID: PMC8599371. DOI: https://doi.org/10.1007/s00424-021-02623-1
Dowling P, Gargan S, Swandulla D, Ohlendieck K. Proteomic profiling of impaired excitation-contraction coupling and abnormal calcium handling in muscular dystrophy. Proteomics. 2022 Dec;22(23-24):e2200003. Epub 2022 Aug 8. PMID: 35902360; PMCID: PMC10078611. DOI: https://doi.org/10.1002/pmic.202200003
Murphy S, Zweyer M, Henry M, Meleady P, Mundegar RR, Swandulla D, Ohlendieck K. Proteomic profiling of liver tissue from the mdx-4cv mouse model of Duchenne muscular dystrophy. Clin Proteomics. 2018 Oct 29;15:34. PMID: 30386187; PMCID: PMC6205794. DOI: https://doi.org/10.1186/s12014-018-9212-2
Dowling P, Zweyer M, Raucamp M, Henry M, Meleady P, Swandulla D, Ohlendieck K. Proteomic and cell biological profiling of the renal phenotype of the mdx-4cv mouse model of Duchenne muscular dystrophy. Eur J Cell Biol. 2020 Jan;99(1):151059. Epub 2019 Nov 18. PMID: 31776009. DOI: https://doi.org/10.1016/j.ejcb.2019.151059
Dowling P, Gargan S, Zweyer M, Sabir H, Henry M, Meleady P, Swandulla D, Ohlendieck K. Proteomic profiling of the interface between the stomach wall and the pancreas in dystrophinopathy. Eur J Transl Myol. 2021 Mar 26;31(1):9627. PMID: 33709651; PMCID: PMC8056161. DOI: https://doi.org/10.4081/ejtm.2020.9627
Murphy S, Zweyer M, Henry M, Meleady P, Mundegar RR, Swandulla D, Ohlendieck K. Label-free mass spectrometric analysis reveals complex changes in the brain proteome from the mdx-4cv mouse model of Duchenne muscular dystrophy. Clin Proteomics. 2015 Nov 23;12:27. PMID: 26604869; PMCID: PMC4657206. DOI: https://doi.org/10.1186/s12014-015-9099-0
Murphy S, Zweyer M, Raucamp M, Henry M, Meleady P, Swandulla D, Ohlendieck K. Proteomic profiling of the mouse diaphragm and refined mass spectrometric analysis of the dystrophic phenotype. J Muscle Res Cell Motil. 2019 Mar;40(1):9-28. Epub 2019 Mar 19. PMID: 30888583. DOI: https://doi.org/10.1007/s10974-019-09507-z
Liao CY, Kennedy BK. Mouse models and aging: longevity and progeria. Curr Top Dev Biol. 2014;109:249-85. PMID: 24947239. DOI: https://doi.org/10.1016/B978-0-12-397920-9.00003-2
Dowling P, Gargan S, Zweyer M, Swandulla D, Ohlendieck K. Extracellular Matrix Proteomics: The mdx-4cv Mouse Diaphragm as a Surrogate for Studying Myofibrosis in Dystrophinopathy. Biomolecules. 2023 Jul;13,1108. PMID: 37509144. DOI: https://doi.org/10.3390/biom13071108
Ackert-Bicknell CL, Anderson LC, Sheehan S, Hill WG, Chang B, Churchill GA, Chesler EJ, Korstanje R, Peters LL. Aging Research Using Mouse Models. Curr Protoc Mouse Biol. 2015 Jun 1;5(2):95-133. PMID: 26069080; PMCID: PMC4590775. DOI: https://doi.org/10.1002/9780470942390.mo140195
Dowling P, Gargan S, Zweyer M, Henry M, Meleady P, Swandulla D, Ohlendieck K. Proteome-wide Changes in the mdx-4cv Spleen due to Pathophysiological Cross Talk with Dystrophin-Deficient Skeletal Muscle. iScience. 2020 Aug 26;23(9):101500. Epub ahead of print. PMID: 32916630; PMCID: PMC7490529. DOI: https://doi.org/10.1016/j.isci.2020.101500
Dowling P, Gargan S, Zweyer M, Henry M, Meleady P, Swandulla D, Ohlendieck K. Protocol for the Bottom-Up Proteomic Analysis of Mouse Spleen. STAR Protoc. 2020 Dec 3;1(3):100196. PMID: 33377090; PMCID: PMC7757555. DOI: https://doi.org/10.1016/j.xpro.2020.100196
Wiśniewski JR. Filter-Aided Sample Preparation for Proteome Analysis. Methods Mol Biol. 2018;1841:3-10. PMID: 30259475. DOI: https://doi.org/10.1007/978-1-4939-8695-8_1
Mi H, Ebert D, Muruganujan A, Mills C, Albou LP, Mushayamaha T, Thomas PD. PANTHER version 16: a revised family classification, tree-based classification tool, enhancer regions and extensive API. Nucleic Acids Res. 2021 Jan 8;49(D1):D394-D403. PMID: 33290554; PMCID: PMC7778891. DOI: https://doi.org/10.1093/nar/gkaa1106
Taraslia VK, Kouskoukis A, Anagnostopoulos AK, Stravopodis DJ, Margaritis LH, Tsangaris GT. Proteomic analysis of normal murine brain parts. Cancer Genomics Proteomics. 2013 May-Jun;10(3):125-54. PMID: 23741028.
Ghanavatinejad F, Fard Tabrizi ZP, Omidghaemi S, Sharifi E, Møller SG, Jami MS. Protein biomarkers of neural system. J Otol. 2019 Sep;14(3):77-88. DOI: https://doi.org/10.1016/j.joto.2019.03.001
Korovesi AG, Anagnostopoulos AK, Pierros V, Stravopodis DJ, Tsangaris GT. Normal Mouse Brain Proteome II: Analysis of Brain Regions by High-resolution Mass Spectrometry. Cancer Genomics Proteomics. 2020 Nov-Dec;17(6):757-767. DOI: https://doi.org/10.21873/cgp.20230
Ishihama Y, Sato T, Tabata T, Miyamoto N, Sagane K, Nagasu T, Oda Y. Quantitative mouse brain proteomics using culture-derived isotope tags as internal standards. Nat Biotechnol. 2005 May;23(5):617-21. Epub 2005 Apr 17. PMID: 15834404. DOI: https://doi.org/10.1038/nbt1086
Seefeldt I, Nebrich G, Römer I, Mao L, Klose J. Evaluation of 2-DE protein patterns from pre- and postnatal stages of the mouse brain. Proteomics. 2006 Sep;6(18):4932-9. PMID: 16912972. DOI: https://doi.org/10.1002/pmic.200600188
Föcking M, Boersema PJ, O'Donoghue N, Lubec G, Pennington SR, Cotter DR, Dunn MJ. 2-D DIGE as a quantitative tool for investigating the HUPO Brain Proteome Project mouse series. Proteomics. 2006 Sep;6(18):4914-31. PMID: 16927420. DOI: https://doi.org/10.1002/pmic.200600269
Sato T, Ishihama Y, Oda Y. Quantitative proteomics of mouse brain and specific protein-interaction studies using stable isotope labeling. Methods Mol Biol. 2007;359:53-70. PMID: 17484110. DOI: https://doi.org/10.1007/978-1-59745-255-7_4
Wang J, Gu Y, Wang L, Hang X, Gao Y, Wang H, Zhang C. HUPO BPP pilot study: a proteomics analysis of the mouse brain of different developmental stages. Proteomics. 2007 Nov;7(21):4008-15. PMID: 17922513. DOI: https://doi.org/10.1002/pmic.200700341
Jung SY, Choi JM, Rousseaux MW, Malovannaya A, Kim JJ, Kutzera J, Wang Y, Huang Y, Zhu W, Maity S, Zoghbi HY, Qin J. An Anatomically Resolved Mouse Brain Proteome Reveals Parkinson Disease-relevant Pathways. Mol Cell Proteomics. 2017 Apr;16(4):581-593. Epub 2017 Feb 2. PMID: 28153913; PMCID: PMC5383780. DOI: https://doi.org/10.1074/mcp.M116.061440
Zhu F, Cizeron M, Qiu Z, Benavides-Piccione R, Kopanitsa MV, Skene NG, Koniaris B, DeFelipe J, Fransén E, Komiyama NH, Grant SGN. Architecture of the Mouse Brain Synaptome. Neuron. 2018 Aug 22;99(4):781-799.e10. Epub 2018 Aug 2. PMID: 30078578; PMCID: PMC6117470. DOI: https://doi.org/10.1016/j.neuron.2018.07.007
Davis RG, Park HM, Kim K, Greer JB, Fellers RT, LeDuc RD, Romanova EV, Rubakhin SS, Zombeck JA, Wu C, Yau PM, Gao P, van Nispen AJ, Patrie SM, Thomas PM, Sweedler JV, Rhodes JS, Kelleher NL. Top-Down Proteomics Enables Comparative Analysis of Brain Proteoforms Between Mouse Strains. Anal Chem. 2018 Mar 20;90(6):3802-3810. Epub 2018 Feb 26. PMID: 29481055; PMCID: PMC5861018. DOI: https://doi.org/10.1021/acs.analchem.7b04108
Park HM, Satta R, Davis RG, Goo YA, LeDuc RD, Fellers RT, Greer JB, Romanova EV, Rubakhin SS, Tai R, Thomas PM, Sweedler JV, Kelleher NL, Patrie SM, Lasek AW. Multidimensional Top-Down Proteomics of Brain-Region-Specific Mouse Brain Proteoforms Responsive to Cocaine and Estradiol. J Proteome Res. 2019 Nov 1;18(11):3999-4012. Epub 2019 Oct 2. PMID: 31550894; PMCID: PMC6917473. DOI: https://doi.org/10.1021/acs.jproteome.9b00481
Sun X, Sun H, Han X, Chen PC, Jiao Y, Wu Z, Zhang X, Wang Z, Niu M, Yu K, Liu D, Dey KK, Mancieri A, Fu Y, Cho JH, Li Y, Poudel S, Branon TC, Ting AY, Peng J. Deep Single-Cell-Type Proteome Profiling of Mouse Brain by Nonsurgical AAV-Mediated Proximity Labeling. Anal Chem. 2022 Apr 5;94(13):5325-5334. Epub 2022 Mar 22. PMID: 35315655; PMCID: PMC9350993. DOI: https://doi.org/10.1021/acs.analchem.1c05212
Rayaprolu S, Bitarafan S, Santiago JV, Betarbet R, Sunna S, Cheng L, Xiao H, Nelson RS, Kumar P, Bagchi P, Duong DM, Goettemoeller AM, Oláh VJ, Rowan M, Levey AI, Wood LB, Seyfried NT, Rangaraju S. Cell type-specific biotin labeling in vivo resolves regional neuronal and astrocyte proteomic differences in mouse brain. Nat Commun. 2022 May 25;13(1):2927. PMID: 35614064; PMCID: PMC9132937. DOI: https://doi.org/10.1038/s41467-022-30623-x
Gomez AM, Traunmüller L, Scheiffele P. Neurexins: molecular codes for shaping neuronal synapses. Nat Rev Neurosci. 2021 Mar;22(3):137-151. Epub 2021 Jan 8. PMID: 33420412; PMCID: PMC7612283. DOI: https://doi.org/10.1038/s41583-020-00415-7
Liu X, Hua F, Yang D, Lin Y, Zhang L, Ying J, Sheng H, Wang X. Roles of neuroligins in central nervous system development: focus on glial neuroligins and neuron neuroligins. J Transl Med. 2022 Sep 10;20(1):418. PMID: 36088343; PMCID: PMC9463862. DOI: https://doi.org/10.1186/s12967-022-03625-y
Rao A, Harms KJ, Craig AM. Neuroligation: building synapses around the neurexin-neuroligin link. Nat Neurosci. 2000 Aug;3(8):747-9. PMID: 10903560. DOI: https://doi.org/10.1038/77636
Montenegro-Venegas C, Guhathakurta D, Pina-Fernandez E, Andres-Alonso M, Plattner F, Gundelfinger ED, Fejtova A. Bassoon controls synaptic vesicle release via regulation of presynaptic phosphorylation and cAMP. EMBO Rep. 2022 Aug 3;23(8):e53659. Epub 2022 Jun 29. PMID: 35766170; PMCID: PMC9346490. DOI: https://doi.org/10.15252/embr.202153659
Smith LM, Kelleher NL; Consortium for Top Down Proteomics. Proteoform: a single term describing protein complexity. Nat Methods. 2013 Mar;10(3):186-7. PMID: 23443629; PMCID: PMC4114032. DOI: https://doi.org/10.1038/nmeth.2369
Forgrave LM, Wang M, Yang D, DeMarco ML. Proteoforms and their expanding role in laboratory medicine. Pract Lab Med. 2021 Nov 27;28:e00260. PMID: 34950758; PMCID: PMC8672040. DOI: https://doi.org/10.1016/j.plabm.2021.e00260
Witzmann FA, Arnold RJ, Bai F, Hrncirova P, Kimpel MW, Mechref YS, McBride WJ, Novotny MV, Pedrick NM, Ringham HN, Simon JR. A proteomic survey of rat cerebral cortical synaptosomes. Proteomics. 2005 May;5(8):2177-201. PMID: 15852343; PMCID: PMC1472619. DOI: https://doi.org/10.1002/pmic.200401102
Errante L, Tang D, Gardon M, Sekerkova G, Mugnaini E, Shaw G. The intermediate filament protein peripherin is a marker for cerebellar climbing fibres. J Neurocytol. 1998 Feb;27(2):69-84. DOI: https://doi.org/10.1023/A:1006991104595
Figueiro-Silva J, Gruart A, Clayton KB, Podlesniy P, Abad MA, Gasull X, Delgado-García JM, Trullas R. Neuronal pentraxin 1 negatively regulates excitatory synapse density and synaptic plasticity. J Neurosci. 2015 Apr 8;35(14):5504-21. PMID: 25855168; PMCID: PMC6605318. DOI: https://doi.org/10.1523/JNEUROSCI.2548-14.2015
Dibattista M, Al Koborssy D, Genovese F, Reisert J. The functional relevance of olfactory marker protein in the vertebrate olfactory system: a never-ending story. Cell Tissue Res. 2021 Jan;383(1):409-427. Epub 2021 Jan 15. PMID: 33447880; PMCID: PMC7878404. DOI: https://doi.org/10.1007/s00441-020-03349-9
Yu C, Kastin AJ, Ding Y, Pan W. Gamma glutamyl transpeptidase is a dynamic indicator of endothelial response to stroke. Exp Neurol. 2007 Jan;203(1):116-22. Epub 2006 Sep 14. PMID: 16973162. DOI: https://doi.org/10.1016/j.expneurol.2006.07.023
Bamford RA, Widagdo J, Takamura N, Eve M, Anggono V, Oguro-Ando A. The Interaction Between Contactin and Amyloid Precursor Protein and Its Role in Alzheimer's Disease. Neuroscience. 2020 Jan 1;424:184-202. Epub 2019 Nov 6. PMID: 31705890. DOI: https://doi.org/10.1016/j.neuroscience.2019.10.006
Li D, Liu X, Liu T, Liu H, Tong L, Jia S, Wang YF. Neurochemical regulation of the expression and function of glial fibrillary acidic protein in astrocytes. Glia. 2020 May;68(5):878-897. Epub 2019 Oct 18. PMID: 31626364. DOI: https://doi.org/10.1002/glia.23734
How to Cite

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
PAGEPress has chosen to apply the Creative Commons Attribution NonCommercial 4.0 International License (CC BY-NC 4.0) to all manuscripts to be published.

